Beginning Bioinformatics: What's a terminal? What's the command line?
| Updated:

Hi! I'm Ryan Moore, PhD bioinformatics data scientist, functional programming fan, and sports nerd. Contact me on LinkedIn!
Installing and running typical bioinformatics programs requires a lot of background knowledge. For beginners, terms like “command line,” “terminal,” “changing directories,” and “archive file” might be unfamiliar. Even instructions to type make can be confusing. There is a lot of prerequisite knowledge needed to get started with installing and using bioinformatics software.
So, let’s start with the basics: terminals and the the command line.
Graphical vs. command line interfaces
You’re probably reading this blog in a web-browser. Whether it’s on a phone or on a laptop, your web-browser is a graphical user interface (GUI). We interact with GUIs by clicking around with the mouse, or if we’re using a mobile device, by tapping and swiping on the screen. Most of the programs we use on our phones and computers are GUIs. Finder on a Mac and Windows Explorer (or File Explorer) on a PC are GUIs that let you browse and manage files on your computer. Chrome, Safari, and Internet Explorer are GUIs for browsing the web. So a GUI is a program with a graphical user interface, and make up the majority of the programs you probably use on a daily basis.
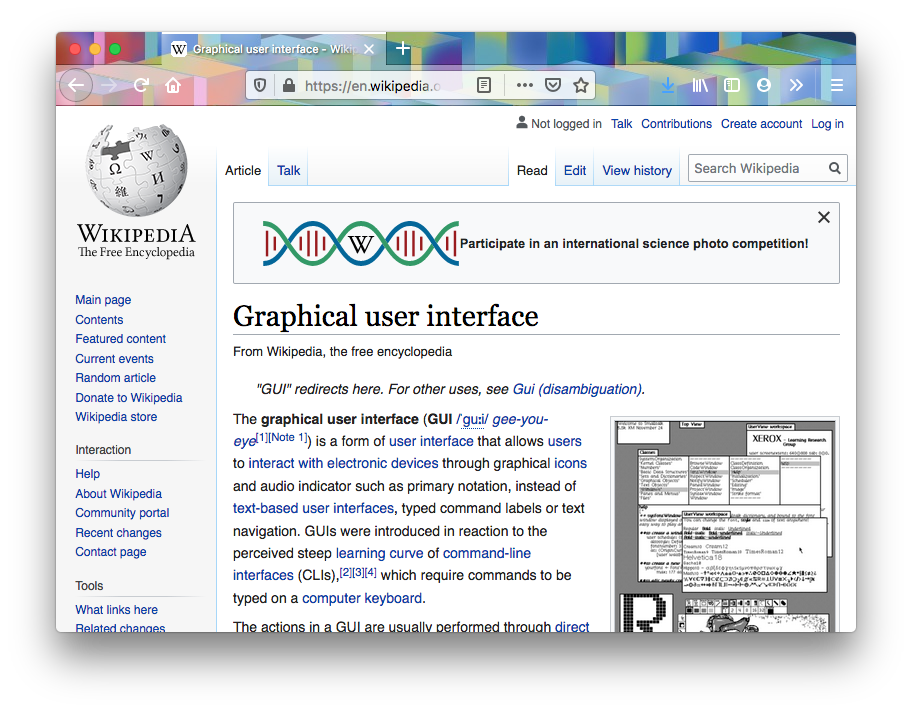
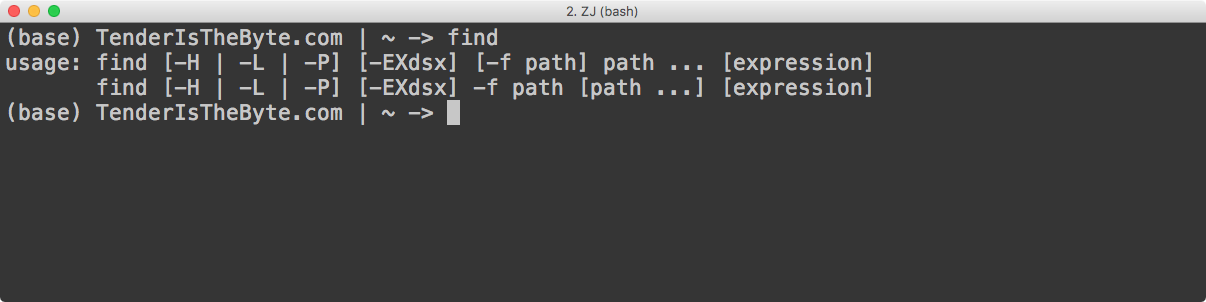
Compare this to programs with so-called command-line interfaces (CLI). Rather than pointing and clicking, you interact with these programs by typing things at the command line, generally through a terminal. One example a program with a command-line interface is find, which finds files based on some user-specified criteria. Most bioinformatics programs don’t have graphical user interfaces. If you want to learn to do bioinformatics, you’re almost certainly going to have to get comfortable with the command line.
Some programs have both a graphical and a command line interface, like Cytoscape, a program for visualizing networks. Why have both? Well, there are some tasks that are easier to accomplish using a graphical user interface and some that are easier with a command line interface. For example, if you need to explore your network–color it, change the size of nodes and edges, make it look nice and pretty–you’re probably going to want to use the Cytoscape GUI for that. If you have an algorithm or process that you want to apply to hundreds of networks, then you’re definitely going to want to use the command line interface instead.
The terminal and the command line
A terminal is a text-based interface to your computer. Depending on who you’re talking to, you might hear the terminal called a couple of different things. The console, the shell, the command prompt–whatever they call it, people are generally talking about the same thing: a place where you enter commands and interact with command-line programs. Of course, all of these terms have more precise definitions, (just ask a systems admin!). For now though, let’s just agree to call it the terminal and not worry too much about it.
As I mentioned earlier, you control a program with a command line interface by typing commands into a terminal. Here is an example of how you might use the find command:
find . -name '*.txt'Let’s talk about this just a bit. The first thing there is the word find. find is the name of the command we’re running. (If you’re reading program documentation or a blog and you see a word in font that looks like this, then it generally means it’s either a command, something you’re typing at the terminal, or some snippet of code.) Next are the arguments that we pass to the find command/program. Arguments let us modify the behavior of a command or program. In this case, the . tells find look in the current directory, and -name '*.txt' bit tells find to look for files that end with .txt. Don’t worry too much if that doesn’t make sense right now. We’ll get into the details of actually running command line programs in a different post. For now, just know that command line programs are those that you control by typing commands and arguments into the termial.
Let me just mention one more thing. If you’re reading program documentation or tutorials about the command line, you might see commands that look like they start with a $ character like this:
$ find . -name '*.txt'The $ character isn’t actually part of the command. Some authors will put it in front of the actual command to represent the command prompt (the place where you’re actually typing in the terminal). It’s just there to make it clearer that what you see is a command that you should type into a terminal.
How do I get a terminal?
If you’re on a Mac, you should have a program called Terminal already installed. To open it, click on the Launchpad and type Terminal into the search box and double click on its icon. iTerm2 is another popular terminal emulator for Macs. If you’re using Linux, then you’ve got tons of options for terminals as well. Windows is a bit different from the other two, but it does have a terminal. Check out this software repository and this guide for more information on the Windows command prompt. I personally don’t use a PC for work, but many people I know who do use PCs for bioinformatics use Cygwin, which let’s you get a more Linux-y command line experience on your PC.
Wrap up
In this post, we talked about graphical user interfaces versus command line interfaces, what is a terminal, what is the command line and how to actually get a terminal for your computer. This is all foundational stuff that you’ll be getting a lot more experience with as you learn more about bioinformatics. Hopefully, this guide helps clear up any confusion you may have had!
If you want some more hands-on info about command line basics, check out this nice tutorial from Django Girls.
If you enjoyed this post, consider sharing it on Twitter or LinkedIn and subscribing to the RSS feed! If you have questions or comments, you can find me on LinkedIn or send me an email directly.
← Go back